Today, the HUNT 1-4 Study is a database with information on approximately 140,000 people that integrates family data and individual data that can be linked to regional EMRs and national health registries.
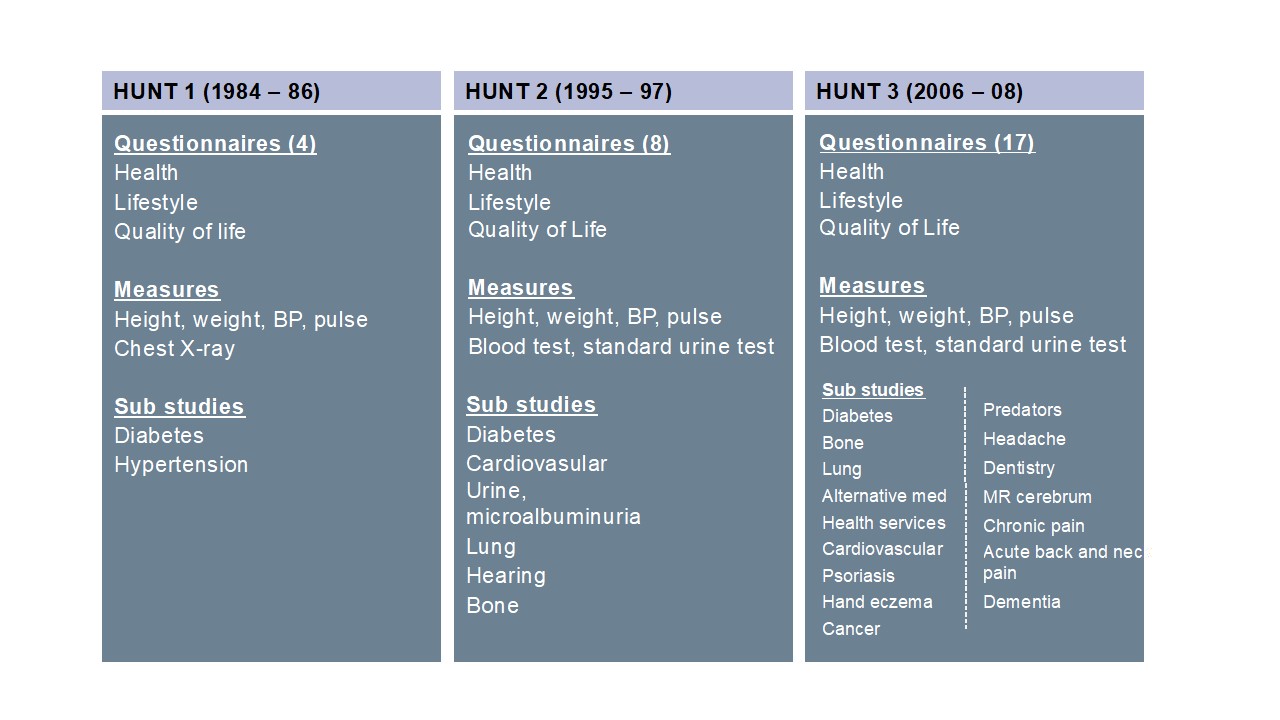
The HUNT4 study (completed in February 2019) was run in a similar way as HUNT3 and comprises questionnaires, interview and clinical exams. New features in HUNT4 include activity sensors and body composition (impedance) as well as a special focus on the elderly above 70 years of age with cognitive and memory testing, gait test and functional MRI on subsets.
Repeated examinations and follow-up of the same population make it possible to ascertain changes in health and vital status at individual and family levels. The HUNT Study is reinforced and supplemented by cross-referencing with structured EMRs data (ICD10 and procedures done at regional hospitals) and separate endpoint registries in diseases such as radial and hip fractures, ischemic heart disease and stroke. The HUNT consent allow cross-linking with registries at the national level (The Cancer Register, The Medical Birth Register). Additionally, Statistics Norway provides necessary information from The Population Census Register and The Family Register to create a genealogical database ("family trees").
A detailed overview and a searchable database of all available variables, with metadata, are available directly from the HUNT databank https://hunt-db.medisin.ntnu.no/hunt-db/#/.
HUNT genetic data
Approximately 88,000 HUNT participants from the HUNT2-4 surveys have available genetic data from array genotyping. DNA was extracted at the HUNT Biobank laboratories (Levanger, Norway), and the genotyping was performed at NTNU Genomic Core Facility (Trondheim, Norway) using Illumina HumanCoreExome arrays. The data has been imputed using 1) sequenced samples (2,200 HUNT samples whole genome sequenced) for joint imputation with the Haplotype Reference Consortium (HRC) imputation panel, and 2) TOPMed imputation panel, and includes around 25 million well-imputed variants. The data is securely stored using the HUNT Cloud.
A good summary of what genetic data other researchers can apply for access to is described in a publication in Cell Genomics 2022 by Brumpton et al. A range of studies of genetic variation on a population level have been performed in HUNT, to a large extent genome-wide association studies (GWAS) and Mendelian Randomization (MR) studies. Compiling a composite score for risk of several diseases based on multiple genes into a polygenic risk score and investigating that score in relation to disease outcomes has also been performed. We refer to this publication for further details.
HUNT Microbiomics
Metagenome sequencing and microbiome profiling has been performed for fecal samples from about 13,000 HUNT4 participants collected as part of the HUNT4 survey. DNA was extracted at the HUNT biobank laboratories (Levanger, Norway) and sent to the Clinical Microbiomics (Copenhagen, Denmark) laboratories, where all non-human DNA was sequenced and analysed.
Sequencing of DNA in the samples has resulted in quantification of the presence of over 6,500 species resulting in quantification of 1) the presence/abundance of over 6,800 microbial species (bacteria, eukaryotes and archaea), of which about 4,900 species were present in at least one participant, 2) their estimated functional potential, 3) the presence/abundance of more than 26,000 viruses, and 4) measures of alpha and beta diversity. The current microbiome data was obtained using the Clinical Microbiomics Human Microbiome Profiler (CHAMP) pipeline and is annotated to the Genome Taxonomy Database (GTDB) nomenclature, while microbiome profiles obtained with the MetaPhlAn 4 pipeline will be available soon.
HUNT Proteomics
HUNT has data generated from blood plasma or serum using platforms from SomaLogic panels and Olink Target panels. The most recent and largest batch of HUNT samples (>2000 samples) with proteomics data are those analysed using the SomaLogic 7000 v4 protein panel in samples from the HUNT3 survey. The project design was case-cohort and the samples were selected based on a focus on cardiovascular disease. The SomaLogic technology is based on the use of modified aptamers, which are short synthetic DNA or RNA molecules that can bind to specific proteins with high affinity and specificity. These aptamers are called SOMAmers and they are designed to recognize and quantify over 7,000 proteins in every blood sample. Different subsets of HUNT samples have been analysed using the panels targeting either 1,000, 3,000 or 5,000 proteins.
There are several subsets generated in past projects (e.g. related to lung cancer, myocardial infarction) that have been analysed using several of the Olink target panels targeting 92 proteins (e.g. cardiovascular or oncology panels). Olink technology is a proteomics platform that uses a method called Proximity Extension Assay (PEA) to measure up to thousands of proteins in a single blood sample with high specificity and sensitivity.
HUNT Metabolomics
Large-scale metabolomic profiling in blood has been performed in samples from >18,000 participants in the HUNT 3 survey, quantifying >240 small-molecular metabolites and lipoprotein subfractions. The samples were analysed by nuclear magnetic resonance (NMR) spectroscopy using the Nightingale quantification assay at the University of Bristol. Additionally, metabolomics profiling of blood samples from 2,400 women in the HUNT2 survey has been performed by NMR spectroscopy using the Bruker BioSpin quantification assays at NTNU.